基于高压液相色谱法通过检测没食子酸测定奇异链霉菌TBGS10的单宁酰基水解酶活性
DOI:
https://doi.org/10.18686/zhfnc.v1i3.102关键词:
奇异链霉菌;单宁酶;高效液相色谱法;没食子酸生产;固态发酵摘要
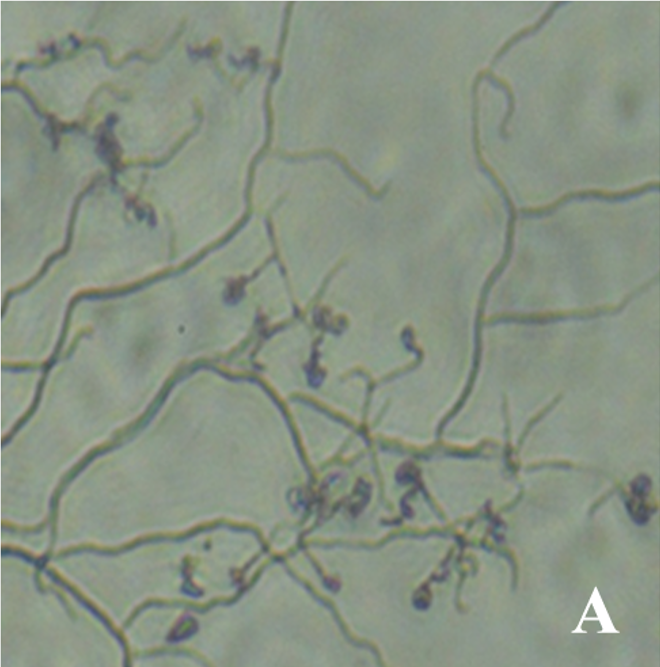
单宁酶是一种水解酶,被称为单宁酰基水解酶,可作用于可水解单宁的酯键并产生没食子酸。单宁酶有多种来源。来自微生物,特别是曲霉属真菌的单宁酶已被用于食品、酿造和制药行业。尽管众所周知放线菌能产生多种工业酶,但放线菌的单宁酰水解酶活性鲜有报道。这篇文章论述了从喀拉拉邦西高止山脉慕那尔的娑罗森林中分离出的链霉菌的单宁酸酶活性。根据形态特征和16s rDNA同源性分析,该分离物被鉴定为奇异链霉菌TBGS10。该分离物在平板试验、深层和固态发酵条件下均表现出良好的单宁酶活性。高压液相色谱法测定了以腰果苹果渣为基质通过固态发酵产生的具有重要工业价值的中间产物没食子酸。提取物观察到没食子酸(gallic acid,GA)含量为142.624 g/mL,保留时间为2.506 min。利用根据NCBI-GenBank中类似链霉菌序列设计的特异引物,对菌株TBGS10的单宁酶基因进行了PCR扩增。
##submission.downloads##
已出版
文章引用
期
栏目
执照
版权声明
CC BY-NC 4.0作者应保留其作品的版权,并授予期刊/出版商首次出版该作品的权利,同时根据以下条款获得许可: 知识共享署名-非商业性4.0国际版(CC BY-NC 4.0)。本许可证允许复制、分发和传播作品,前提是声明了原创作者的正确归属。
参考
1. Chávez-González M, Rodríguez-Durán LV, Balagurusamy N. Biotechnological advances and challenges of tannase: An overview. Food Bioprocess Technology 2012; 5(2): 445–459. doi: 10.1007/s11947-011-0608-5
2. Wu C, Zhang F, Li L, et al. Novel optimization strategy for tannase production through a modified solid-state fermentation system. Biotechnology for Biofuels and Bioproducts 2018; 11: 92. doi: 10.1186/s13068-018-1093-0
3. Jana A, Halder SK, Ghosh K, et al. Tannase immobilization by chitin-alginate based adsorption entrapment technique and its exploitation in fruit juice clarification. Food Bioprocess Technology 2015; 8: 2319–2329. doi: 10.1007/s11947-015-1586-9
4. Beniwal V, Kumar A, Sharma J, Chhokar V. Recent advances in industrial application of tannases: A review. Recent Patents on Biotechnology 2013; 7(3): 228–233. doi: 10.2174/18722083113076660013
5. Dhiman S, Mukherjee G, Singh AK. Recent trends and advancements in microbial tannase-catalyzed biotransformation of tannins: A review. International Microbiology 2018; 21(4): 175–195. doi: 10.1007/s10123-018-0027-9
6. Farha AK, Yang QQ, Kim G, et al. Tannins as an alternative to antibiotics. Food Bioscience 2020; 38: 100751. doi: 10.1016/j.fbio.2020.100751
7. García Méndez MG, Morales Martínez TK, Ascacio Valdés JA, et al. Application of lactic acid bacteria in fermentation processes to obtain tannases using agro-industrial wastes. Fermentation 2021; 7(2): 48. doi: 10.3390/fermentation7020048
8. Barrios-González J. Solid-state fermentation: Physiology of solid medium, its molecular basis and applications. Process Biochemistry 2012; 47(2): 175–185. doi: 10.1016/j.procbio.2011.11.016
9. Aharwar A, Parihar DK. Tannases: Production, properties, applications. Biocatalysis and Agricultural Biotechnology 2018; 15: 322–334. doi: 10.1016/j.bcab.2018.07.005
10. Nishitani Y, Osawa R. A novel colorimetric method to quantify tannase activity of viable bacteria. Journal of Microbiological Methods 2003; 54(2): 281–284. doi: 10.1016/S0167-7012(03)00063-0
11. Girdhari SN, Peshwe SA. Screening of agro-residues for the production of microbial tannase. International Journal of Advanced Research 2017; 5: 625–631. doi: 10.21474/IJAR01/2790
12. Sharma S, Bhat TK, Dawra RK. A spectrophotometric method for assay of tannase using rhodanine. Analytical Biochemistry 2000; 279(1): 85–89. doi: 10.1006/abio.1999.4405
13. Shirling EB, Gottlieb D. Methods for characterization of Streptomyces species. International Journal of Systematic Bacteriology 1966; 16(3): 313–340. doi: 10.1099/00207713-16-3-313
14. Chakrabarti T. Detection of Polyene Class of Antifungal Antibiotics. Actinomycetes-Isolation, Screening, Identification and Gene Cloning in Streptomyces. Laboratory Manual. Institute of Microbial Technology (IMTECH); 1998.
15. Murray MG, Thompson WF. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Research 1980; 8(19): 4321–4326. doi: 10.1093/nar/8.19.4321
16. Zuckerkandl E, Pauling L. Evolutionary divergence and convergence in proteins. In: Bryson V, Vogel HJ (editors). Evolving Genes and Proteins. Academic Press; 1965. pp. 97–166.
17. Tamura K, Stecher G, Peterson D, et al. MEGA6: Molecular evolutionary genetics analysis version 6.0. Molecular Biology and Evolution 2013; 30(12): 2725–2729. doi: 10.1093/molbev/mst197
18. El Sohafy SM, Metwalli AM, Harraz FM, Omar AA. Quantification of flavonoids of Psidium guajava L. preparations by Planar Chromatography (HPTLC). Pharmacognosy Magazine 2009; 4(17): 61–66.
19. Samee W, Vorarat S. Simultaneous determination of gallic acid, catechin, rutin, ellagic acid, and quercetin in flower extracts of Michelia alba, Caesalpinia pulcherrima, and Nelumbo nucifera by HPLC. Thai Pharmaceutical and Health Science Journal 2007; 2: 131–137.
20. Tamura K, Nei M. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Molecular Biology and Evolution 1993; 10(3): 512–526. doi: 10.1093/oxfordjournals.molbev.a040023
21. Roy S, Parvin R, Ghosh S, et al. Occurrence of a novel tannase (tan BLP) in endophytic Streptomyces sp. AL1L from the leaf of Ailanthus excelsa Roxb. 3 Biotech 2018; 8: 33. doi: 10.1007/s13205-017-1055-4
22. Mueller-Harvey I, Parkes RJ. Measurement of volatile fatty acids in pore water from marine sediments by HPLC. Estuarine, Coastal and Shelf Science 1987; 25(5): 567–579. doi: 10.1016/0272-7714(87)90115-6
23. Kumar SS, Sreekumar R, Sabu A. Tannase and its applications in food processing. In: Parameswaran B, Varjani S, Raveendran S (editors). Green Bio-processes. Energy, Environment, and Sustainability. Springer; 2019. pp. 357–381. doi: 10.1007/978-981-13-3263-0_19
24. Sharma S, Gupta MN. Synthesis of antioxidant propyl gallate using tannase from Aspergillus niger van Tiegham in nonaqueous media. Bioorganic & Medicinal Chemistry Letters 2003; 13(3): 395–397. doi: 10.1016/S0960-894X(02)00977-0
25. de Lima JS, Cruz R, Fonseca JC, et al. Production, characterization of tannase from Penicillium montanense URM 6286 under SSF using agroindustrial wastes, and application in the clarification of grape juice (Vitis vinifera L.). The Scientific World Journal 2014; 2014: 182025. doi: 10.1155/2014/182025
26. Neethu RS, Pradeep NS. Isolation and characterization of potential tannase-producing fungi from mangroves and tanneries. Indian Journal of Applied Microbiology 2018; 21(3): 1–13.
27. Tripathi AD, Lakshmi B. Statistical optimization of extracellular tannase production by Streptomyces sp. AT 13 using response surface methodology and Plackett-Burmen design. Bioscience Biotechnology Research Communications 2018; 11(4): 691–698. doi: 10.21786/bbrc/11.4/21
28. Böer E, Bode R, Mock HP, et al. Atan1p—An extracellular tannase from the dimorphic yeast Arxula adeninivorans: Molecular cloning of the ATAN1 gene and characterization of the recombinant enzyme. Yeast 2009; 26(6): 323–337. doi: 10.1002/yea.1669
29. Lekshmi R, Nisha SA, Vasan PT, Kaleeswaran B. A comprehensive review on tannase: Microbes associated production of tannase exploiting tannin rich agro-industrial wastes with special reference to its potential environmental and industrial applications. Environmental Research 2021; 201: 111625. doi: 10.1016/j.envres.2021.111625
30. Podrigues THS, Dantas MAA, Pinto GAS, Gonçalves LRB. Tannase production by solid state fermentation of cashew apple bagasse. In: Mielenz JR, Klasson KT, Adney WS, McMillan JD (editors). Applied Biochemistry and Biotecnology. ABAB Symposium. Humana Press; 2007. pp. 675–688. doi: 10.1007/978-1-60327-181-3_55
31. Kar B, Banerjee R, Bhattacharyya BC. Effect of additives on the behavioural properties of tannin acyl hydrolase. Process Biochemistry 2003; 38(9): 1285–1293. doi: 10.1016/S0032-9592(02)00329-1
32. Pinto GAS, Leite SGF, Terzi SC, Couri S. Selection of tannase-producing Aspergillus niger strains. Brazilian Journal of Microbiology 2001; 32: 24–26. doi: 10.1590/S1517-83822001000100006
33. Seth M, Chand S. Biosynthesis of tannase and hydrolysis of tannins to gallic acid by Aspergillus awamori—Optimisation of process parameters. Process Biochemistry 2000; 36(1–2): 39–44. doi: 10.1016/S0032-9592(00)00179-5
34. Deschamps AM, Lebeault JM. Production of gallic acid from tara tannin by bacterial strains. Biotechnology. Letters 1984; 6: 237–242. doi: 10.1007/BF00140043
35. Nalan Yılmaz S, Erdoğan Ç, Merih K, Muzaffer T. A method for the determination of tannase activity based on gallic acid measurement by high-performance liquid chromatography (HPLC). African Journal of Microbiology Research 2011; 5(2): 158–163. doi: 10.5897/AJMR10.548


